research
The Armbrust group uses lab- and field-based approaches to understand the complexities surrounding phytoplankton, their environment and their interactions with other microbes. We work at the cellular, population, and community scale to understand how these organisms shape and are shaped by environmental conditions.
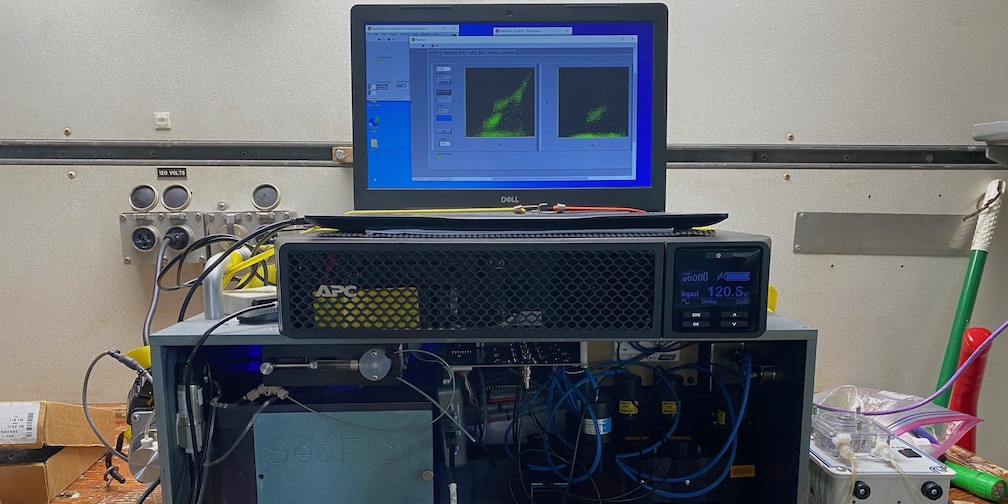
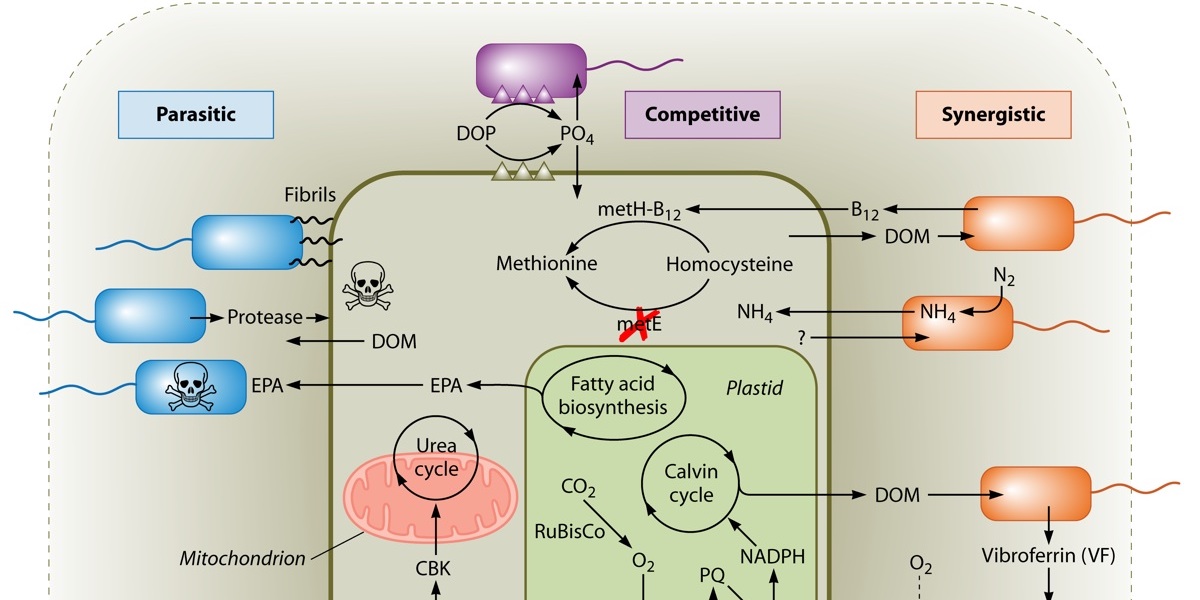
In the lab, we use molecular, physiological, and chemical measurements to examine intra- and inter-species relationships among diatoms, bacteria and archaea. Complete genomes and whole-cell transcriptomes allow us to interrogate diatoms as they respond to nutrient limitation, interact with other microbes, and adapt to projected environmental changes such as increased carbon dioxide levels. We also develop a suite of bioinformatics and flow cytometric software to address current challenges in large-scale data acquisition and analysis. These data encompass DNA/transcript DNA sequence data and underway analyses of flow cytometric phytoplankton distributions.
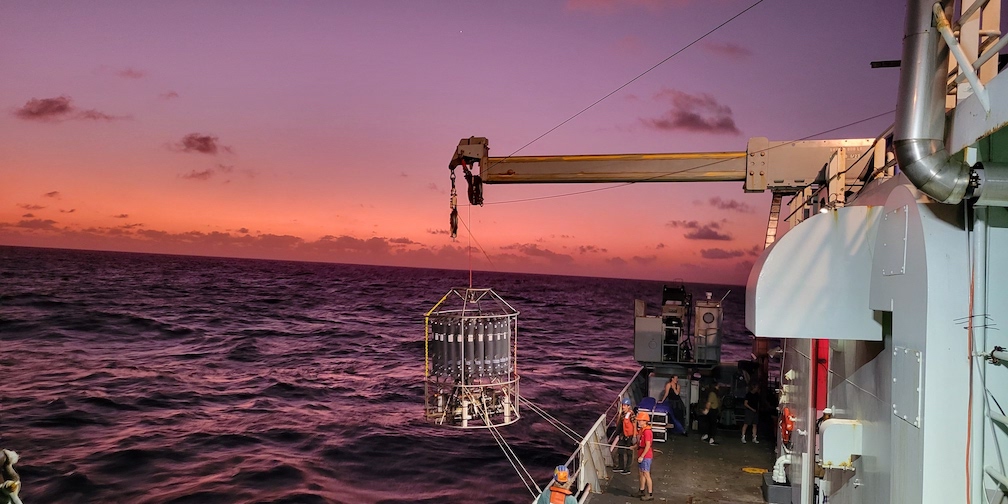
In the field, we collect temporal and spatial samples across environmental gradients to better understand the impacts of community-wide interactions on biogeochemical cycles. Metagenomes, metatranscriptomes and our custom-made underway flow cytometer (SeaFlow) enable high-resolution insight into the abundance and distribution of these microscopic organisms. Measurements are coupled to classic oceanographic parameters (e.g. biogenic silica, nutrients, chlorophyll) to help us assess the biogeochemical ramifications of the impact of diatoms and their interactions with others on the marine environment.